After a few weeks of doing cryogenic cathodoluminescence spectroscopy on some of my samples, I have gobs and gobs of spectra to look over, and the task is a bit daunting. Oh sure, you can do so on the computer in many different ways, but often, I need to see my data on the page (old school) before I can really sort through it. Sometimes even then there’s just too much of it and playing with the data in MatLab all together is critical.
Here’s what I came up with to help output my data very quickly into a printable document that included numerous graphs. First, the primary goal of this quick method is to be quick- to get tons of graphics (of same proportions) into a doc for printing or perusing. Second- it should be relatively minimal typing, if possible. [We all know we can do it by hand 100x; while grad-student pay rate is low, there's gotta be a better way.]
- Get a directory listing for all the items wanted to be included and dump this in a text file. (ls *.pdf > filelist.tex)
- Create a main.tex file which includes the code we’ll need to do this fast. [You can reuse this file for other directories of graphics needed to be printed.] My example uses the following:
\documentclass[%
,secnumarabic%
,amssymb, amsmath,nobibnotes, aps, prl,superscriptaddress,letterpaper]{revtex4}
\usepackage[pdftex]{color,graphicx,rotating}
\setkeys{Gin}{width=0.85\columnwidth}
%Simple way to call images and add filenames to captions - for lots of data.
\newcommand{\dataimg}[1]{
\includegraphics[width=3in]{#1}
Filename: #1
}
\begin{document}
\title{CL Results\\
\textit{Internal document not for distribution.}}
\date{\today}
\author{Allen Hall}
\maketitle
\include{filenames}
\printfigures
\end{document}
The important code is the “\include{}” line and also the “\newcommand{\dataimg}…” section. This is what is going to do all the work for us.
- Now, we need to take your filelist and add at the beginning of each line and end of each line the following:
\dataimg{
and at the end:
}
One way to do this simply is with a command line gawk command:
Terminal Prompt> ls |awk '{ print "\\dataimg{"$0"}" }'
- So, now each line looks like: \dataimg{filename1.pdf}
- Once that is done, you can run the LaTeX compile, and you’ll have your file of graphs! That’s a heck of a lot easier than writing each line out by hand. [Use a program like TextMate or Gawk etc. to append and prepend each line with the necessary call.]
The benefit of the \newcommand is that it fills in the needed formatting for each graphics file, and attaches the filename for each graphic beside the graphics file itself. You can make it prettier, I’m sure, but this is what I was able to do in a very short time frame.
There are many ways to accomplish this little task, you could use Gawk itself to write the latex file for you, I’ve seen some do makefiles to do this type of thing, or perl, or bash shell scripting etc. But, the critical part is to leverage the computer to output a latex file for typsetting and save yourself some time.
In the time I’ve been doing my research work at the Univ. of IL, I’ve come across a number of graphs from various past researchers, older papers, stuck on the side of machines (calibration curves), and even hand-drawn or chart-recorder graphs in my numerous projects. The only major problem with those graphs I’ve found is that they aren’t in a digital form for further use with other data (instrument response functions) or to include in your own work as a reference. So, what to do?
Well, there’s an easy solution. It’s not the perfect solution, as it’s a bit slow, I’ll get to that in a second, but it’s a great solution to the problem, and has worked for me a number of times now. To top it off, it’s open-source, donation-ware, and cross-platform: Engauge Digitizer (see post at LifeHacker.com). Don’t let the website and lack of recent updates deter you. Tools that can do what Engauge does are few and far between. So, it is definitely worth a try. Here’s an example of how I’ve used it just the other day (prompting this post- I’ve used it for years now, but the recent use reminded me I should share it with others). [click "More" to see an example use and learn more]
Read More »
By Allen
|
Also posted in DataVisualization, Materials Science and Engineering, Matlab, OpenSource, Research Work, Software, Thesis Writing
|
Tagged Cross Platform, Data Mining, How To, Linux, Mac OsX, Materials Science and Engineering, Open Source, Software, Thesis Writing, Windows
|
For those friends of mine currently attempting to work in LaTeX to code Materials Science and Engineering related tidbits…
Here’s a quick tip. Tired of typing all those crazy math commands for your material’s name? Simply use a new command. Here’s are a couple examples…
%Simpler way of writing CUINSE2:
\newcommand{\cis}{CuInSe$_{2}$}
%Simpler way of writing CUINGASE2:
\newcommand{\cigs}{CuIn$_{(1-x)}$Ga$_{x}$Se$_{2}$}
%Simpler way of writing CUGASE2:
\newcommand{\cgs}{CuGaSe$_{2}$}
To see how this looks in compiled LaTeX output and another example, click “More”…
Read More »
October 26, 2008 – 9:43 pm
Boston rocked: I’m writing this on a new iPod Touch! Filmetrics was running a drawing for those who brought samples to test! Big thankyou’s go out to Filmetrics and AVS. I will write more about the conference in a series of posts coming soon. I’m hoping everyone had a great week while I was gone.
October 17, 2008 – 3:53 pm
The society formerly known as the American Vacuum Society (AVS) is holding their international conference in Boston, MA next week! And I’ll be there! [Exciting!] If you are going to be there, drop me a line on the blog and we can snag a coffee or beer together.
The programs and cards sent out may have a familiar image! 

My AFM image of the CuInSe2 Bicrystals I grow in our lab won second place in the Art-Zone competition last year! It was great fun! [Thanks goes to AVS for being a fantastic organization!]

Wish me luck with my talk!! 
October 9, 2008 – 3:17 pm
A series of MATLAB functions will follow. Some of these “.m” files call each other, so ensure you have all of them before you start.
The purpose of this file is to load in all the .x00 files in a directory of Omega-Omega-2-Theta slices for reciprocal space mapping of a sample. “OmegAxis” determines the scale for the Omega axis (y) for reciprocal space mapping. [again, this is not yet in Q-space, that will come at a later date]
The first function loads data into your workspace for a directory of .x00 files that have a common base filename. The user needs to supply the first index number and last index number for loading the files. The program then attempts to load “filenameIndex.x00″ in a linear fashion for all .x00 files of that filename indexed in the cd’d directory. [Change directory to the working directory needed before starting this function.]
Read More »
October 8, 2008 – 3:14 pm
A series of MATLAB functions will follow. Some of these “.m” files call each other, so ensure you have all of them before you start.
The purpose of this file is to load in all the .x00 files in a directory of Omega-Omega-2-Theta slices for reciprocal space mapping of a sample. Please note that as of this writing, the data has not yet been converted to Q-space.
The first function loads data into your workspace for a directory of .x00 files that have a common base filename. The user needs to supply the first index number and last index number for loading the files. The program then attempts to load “filenameIndex.x00″ in a linear fashion for all .x00 files of that filename indexed in the cd’d directory. [Change directory to the working directory needed before starting this function.]
Read More »
October 7, 2008 – 3:40 pm
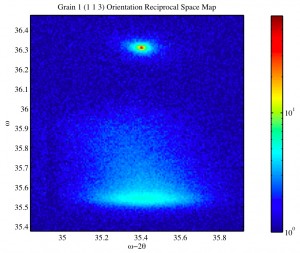
There will be a number of posts following this one that will attempt to recreate reciprocal space mapping in MATLAB. The functions that will be posted in the coming days read in .x00 files from the Philips XPert XRD system and try to recreate the slices of Omega Omega-2Theta scans produced for reciprocal space mapping.
Currently I haven’t changed the data into Q-space, so they will be reported in Omega -v- Omega2Theta space.
An example call to the function is:
DisplayQSpace1('',9,184);
Example output of the function(s):
October 7, 2008 – 3:04 pm
I decided here and there to publish various MATLAB functions I’ve written for my own work. There’s just one caveat- each one of these is a serious work in progress. In fact, they may not progress much more than what is disclosed here since they get the job done for me.
But, I felt that this likely would help a few people who are trying to do the same type of thing, and I’d rather ease those who come after me, and give them a slight step ahead if possible.
For those of you with old version of the Philips XPert XRD software (specifically those who can’t output in XRDML files), I offer a crude import function for simple scans.
Read More »
September 20, 2008 – 10:25 pm
One thing I find extremely helpful in my research, is the ability to download citations for articles my co-workers send. I like to import them into BibDesk and auto-file the journal articles by first-author. Unfortunately, not everyone has Bibtex export abilities. This isn’t too bad, as BibDesk imports most major citation files. One thing that I do abhorr, is when a major company decides not to offer a citation download option! Science Direct is the fellow who gets my wrath this evening. It’s so bad I’ve been looking for other companies who keep track of journals that have downloads for the ones that Science Direct publishes!
I wonder if you pay their absurd journal prices, if you get citation downloads…
For more on Absurd Journal pricing, see: Knuth and “Trapped…”




