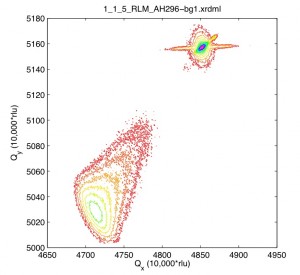
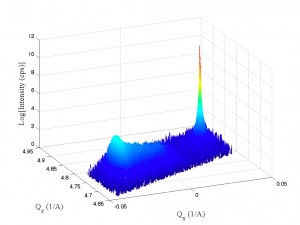
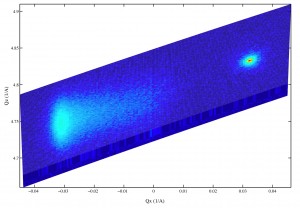
After some new scans, it appears the XRDMLread.m function I talked about is doing a pretty good job of getting the 2-axis scans into MATLAB. I was able to alter the code to accept the standard Epitaxy software’s translation to Q-space. (In Epitaxy the default is R = 0.5 I believe.) So, the following image was imported with XRDMLread.m the plotted with the standard 2-d example from the author’s website. The code was altered to output 10000xRLU units the same as Epitaxy (Panalytical). I haven’t checked all the numbers, but it’s looking ok so far. Unfortunately the color-scale looses it’s meaning as far as intensity is concerned, it appears at first glance.
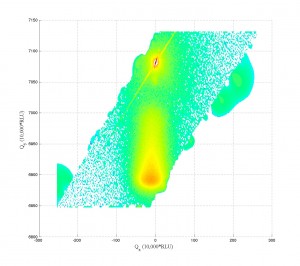
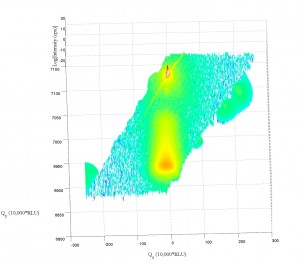
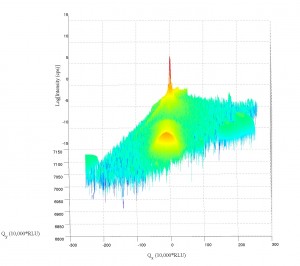
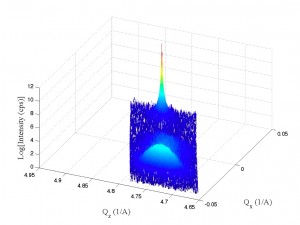
Working a bit with my old q-space map code, I’m able to accomplish the following:
Note the strange love-handles the data gains. I suspect this might be due to the Gridfit function (see MATLAB files repository) I used for regridding the data. Gridfit.m uses an extrapolant method. My suspicion is it is trying to fill out the square of the data matrix and is accomplishing relatively correct values for near-by-data that is outside the scanned range. I’ll try it again with regrid or something similar in the future when I have time.
If anyone knows where the current site for XRDMLread.m is, I’d love to link to it. It appears the site may be down (graduated student I suspect). You can obtain the wonderful XRDMLread.m function and examples on the XRDMLread.m website. For now, I have to wait until I hear from the authors before I can share the file. I also don’t yet trust my icky 3D code, so I prefer not to release that until I have things hashed out. Sorry!
I’m extremely happy that Panalytical has published their XRDML file format and that the makers of XRDML.m have released their .m files for MATLAB. In the past, when Philips had the Xpert systems, the data was stuck for the most part in proprietary data formats. [You could slice the data and output in ascii- but making that work was a pain- which is why I never released that previous code.] I’m much closer to the 3d plotting now, and hope to finish it up before the thesis (my primary work which is not this plotting) is published.
Wishing you luck in you research!